|
Size: 4523
Comment:
|
Size: 4499
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 59: | Line 59: |
| * GMT / 'hsapiens.NAME.gmt' * Dataset 1 / Expression: (OPTIONAL) |
* GMT / '''hsapiens.NAME.gmt''' |
| Line 71: | Line 70: |
| *7. Go to '''View''', and activate '''Show Graphics Details''' | *7. In the menu bar, Go to '''View''', and activate '''Show Graphics Details''' |
Enrichment Map g:Profiler Tutorial
Contents
Outline
This quick tutorial will guide you through the generation of an Enrichment Map for an analysis performed using g:Profiler (Functional Profiling of Gene List from large-scale experiments)
To run this tutorial:
- You need to have Cytoscape installed : minimally 2.6.3 must be installed but preferable to have the latest version of Cytoscape (e.g. version 3.1)
Install the Enrichment Map plugin from the Cytoscape plugin manager. If you install it manually (e.g. if you need to install a new version that doesn't happen to be in the plugin manager yet), then it must be in the Cytoscape-[Version#]/plugins folder --> For Cytoscape 2.8
Install the Enrichment Map App from the Cytoscape App manager in cytoscape 3. --> For Cytoscape 3.
You need to download the test data: gProfilerTutorial.zip
Description of the tutorial files contained in the gProfilerTutorial folder:
- 12hr_topgenes.txt : List of top genes expressed in Estrogen dataset at 12hr - Official Gene Symbol.
- 24hr_topgenes.txt : List of top genes expressed in Estrogen dataset at 24hr - Official Gene Symbol.
- gProfiler_output_12hr.txt : Estrogen treatment - 12hr g:Profiler output results
- gProfiler_output_24hr.txt : Estrogen treatment - 24hr g:Profiler output results
- Estrogen_expression_file.txt: Expression File - Estrogen treatment, Official Gene Name as key.
- hsapiens.NAME.gmt: GMT file (gene-set file) downloaded from g:Profiler website
Instructions
Step 1: Generate g:Profiler output files
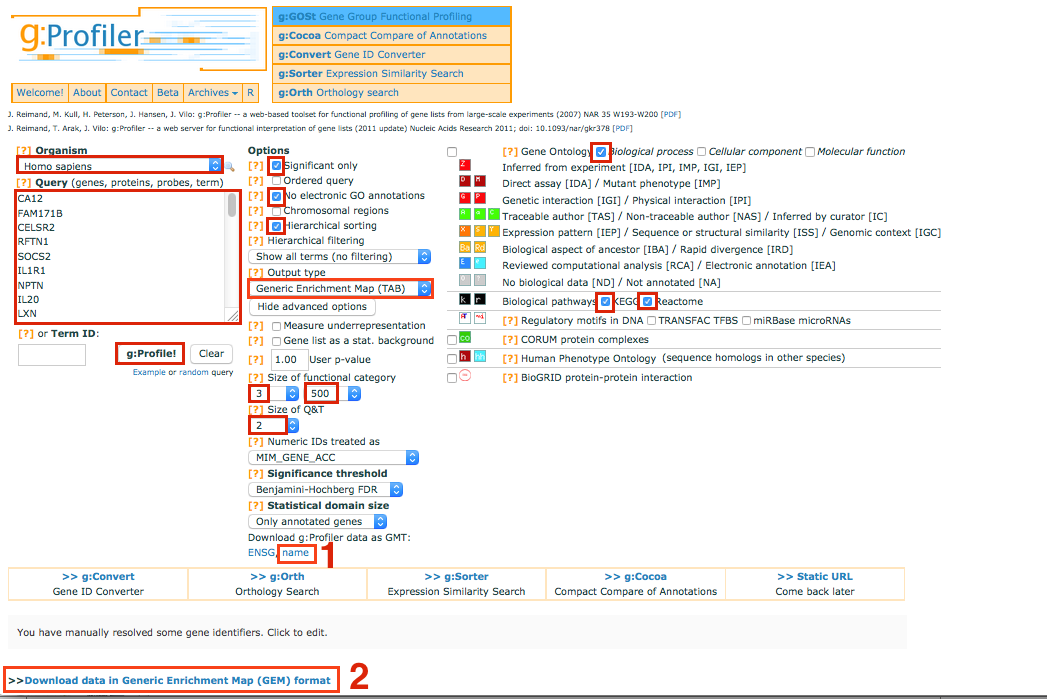
Go to g:Profiler website - http://biit.cs.ut.ee/gprofiler/
Select and copy all genes in the tutorial file 12hr_topgenes.txt in the Query box
In Options, check Significant only, No electronic GO annotations
Set the Output type to Generic EnrichmentMap
Show advanced options
Set Max size of functional category to 500
Set Significance threshold to Benjamini-Hochberg FDR
Click on g:Profile! to run the analysis
Download g:Profiler data as gmt: name
Download the result file: Download data in Generic Enrichment Map (GEM) format
Note: repeat these steps for the 24hrs time-point and the file 24hr_topgenes.txt
Step 2: Generate Enrichment Map with g:Profiler Output
g:Profiler output files gProfiler_EM.zip
1. Open Cytoscape
2. In the menu bar, locate the App tab and then select --> EnrichmentMap --> Create Enrichment Map
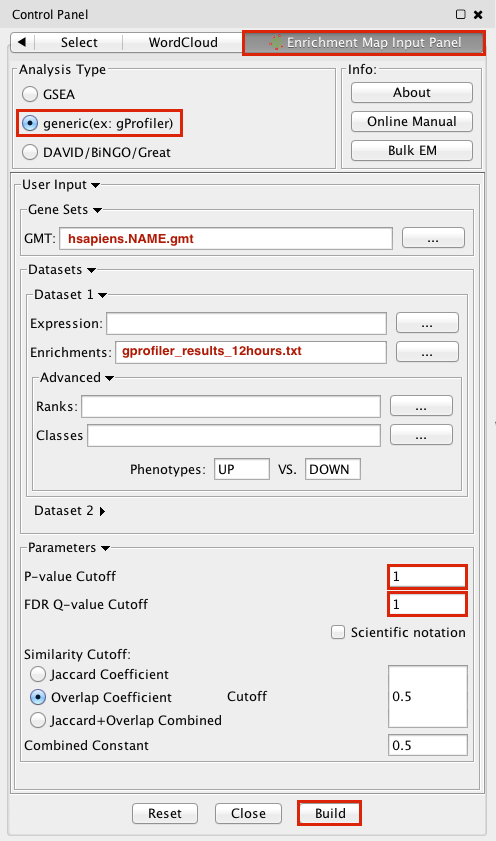
3. Make sure the Analysis Type is set to generic(ex:gProfiler)
4. Please select the following files by clicking on the respective (...) button and selecting the file in the Dialog:
GMT / hsapiens.NAME.gmt
Dataset 1 / Enrichments: gProfiler_results_12hr.txt
5. Tune Parameters
P-value cut-off 1
Q-value cut-off 1
Overlap Coefficient cut-off 0.5
6. Click on the Build radio button at the bottom of the panel to create the Enrichment Map
7. In the menu bar, Go to View, and activate Show Graphics Details
Step 3: Examining Results
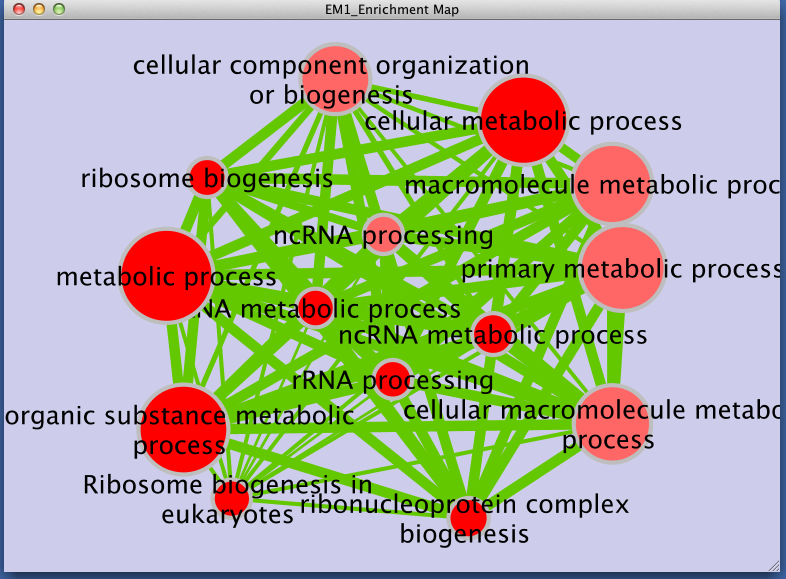
Legend:
- Node (inner circle) size corresponds to the number of genes in dataset 1 within the geneset
- Node border (outer circle) size corresponds to the number of genes in dataset 2 within the geneset
- Colour of the node (inner circle) and border(outer circle) corresponds to the significance of the geneset for dataset 1 and dataset 2, respectively.
- Edge size corresponds to the number of genes that overlap between the two connected genesets. Green edges correspond to both datasets when it is the only colour edge. When there are two different edge colours, green corresponds to dataset 1 and blue corresponds to dataset 2.
